We study the molecular mechanisms that drive microbial pathogenesis. To capture these processes across scales, from atomic structure to cellular function, we use an interdisciplinary approach that combines structural biology (cryo-EM, NMR, X-ray crystallography) with biophysical, cellular, and omics-based techniques. We aim to translate our discoveries into new antibiotics and innovative biotechnological tools.
How Pathogenic Bacteria Build Pili
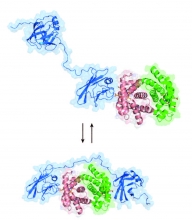
The Clubb Lab uses multidisciplinary approaches to uncover how Gram-positive bacterial pathogens build pili and other surface structures that allow them to attach to tissues, form biofilms, and evade immune defenses. This research provides insight into bacterial virulence and lays the groundwork for developing therapies to combat antibiotic-resistant infections.
...Starving Pathogens: Targeting Heme Acquisition
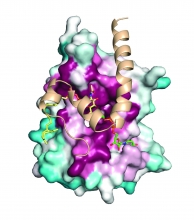
Bacterial pathogens survive in the human host by evading iron sequestration. Our lab studies how they hijack heme from hemoglobin—a process critical for their survival and virulence. Using structural biology, biophysics, proteomics, and protein engineering, we aim to uncover the molecular mechanisms of heme capture and find new ways to block it to combat infectious disease.
...Decoding Microbial Machines
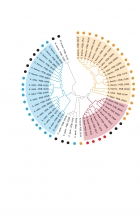
The Clubb Lab at the UCLA–DOE Institute for Genomics and Proteomics studies how bacteria build and control cellulosomes—large enzyme complexes that efficiently break down plant polysaccharides. These systems are central to microbial carbon use and have broad relevance to bioenergy, biotechnology, and the human gut microbiome.
...
-------------------------------------------------------------------------------------------------------------------------
04/25 Esmeralda Corrales joined the lab! Welcome!
-------------------------------------------------------------------------------------------------------------------------
01/25 Nikki Cheung was awarded a Science Olympiad Fellowship!
-------------------------------------------------------------------------------------------------------------------------
08/24 Allen Takayesu won best poster at the Protein Society Meeting!
-------------------------------------------------------------------------------------------------------------------------
07/24 Kylie Cheng was awarded a CMB Fellowship!
-------------------------------------------------------------------------------------------------------------------------
06/24 Kylie Cheng joined the lab! Welcome!
-------------------------------------------------------------------------------------------------------------------------
06/24 Andrew Goring was awarded a UCLA Dissertation Year Fellowship!
-------------------------------------------------------------------------------------------------------------------------
05/24 Nikki Cheung was awarded a UCLA Whitcome Fellowship!
-------------------------------------------------------------------------------------------------------------------------
08/23 Christine Minor was awarded a CMB Fellowship!
-------------------------------------------------------------------------------------------------------------------------
07/23 Andrew Goring was awarded a UCLA Whitcome Fellowship!
-------------------------------------------------------------------------------------------------------------------------
05/23 Reece Pawlacyk joined our research group. Welcome!
-------------------------------------------------------------------------------------------------------------------------
08/22 Dr. Chris Sue was awarded his PhD in biochemistry! Congratulations!
-------------------------------------------------------------------------------------------------------------------------